For many users in the finance, insurance, retail, manufacturing, and even healthcare industries, Power BI has become a staple in their business intelligence plan. From interactive visualizations to advanced data wrangling, Power BI offers a one-stop shop for gaining insights from your data.
However, for any of us that are researchers in the bioinformatics and genomics space, we know that our files can be a bit difficult to work with. From FASTA to BAM, working with files in bioinformatics add a layer of uniqueness that requires some special care.
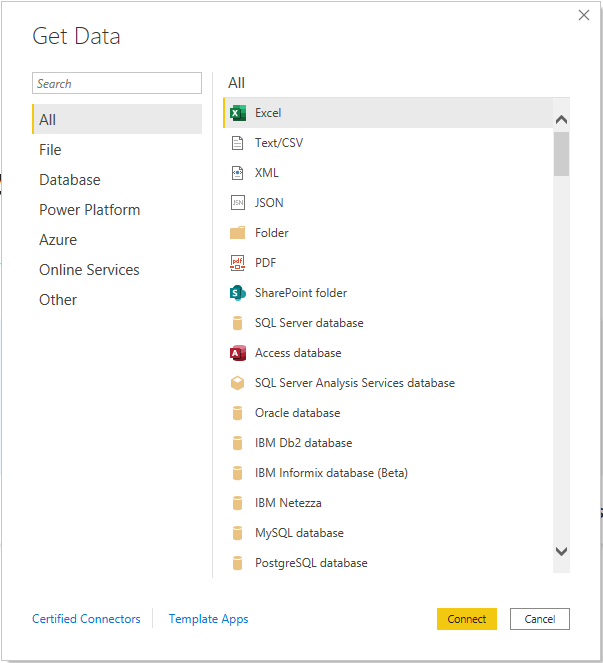
Today, if you take a look at Power BI Desktop’s options for getting data, you’ll see a ton of sources to which you can easily connect. One problem: none of these uniquely help us bioinformaticians.
Putting the “Power” in Power Query
In bioinformatics, there are a plethora of file types for every occasion. Among these are very popular ones such as FASTA (or FASTQ) and BAM and, more recently, GFF3 and BGEN. We can break these data sources down into three main types:
 |
 |
 |
| Text-Based Files
Files that are human readable and can be open using virtually any text editor. |
Binary Files
Files that are serialized and must be read by machines. |
Online Sources
Databases, webpages, or FTP sites on the internet. |
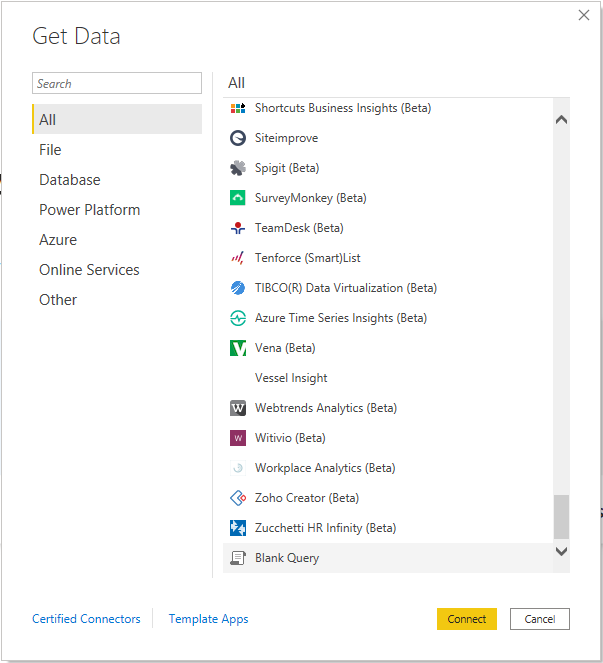
In Power BI, we can take advantage of Power Query to read in data and parse it appropriately. You’ll notice that, while Power BI has tons of connects to everything from CSV files to Spark clusters, there are no built-in connectors for our beloved genomics file types (yet?). So, we’ll have to use the Blank Query editor.
Text Files
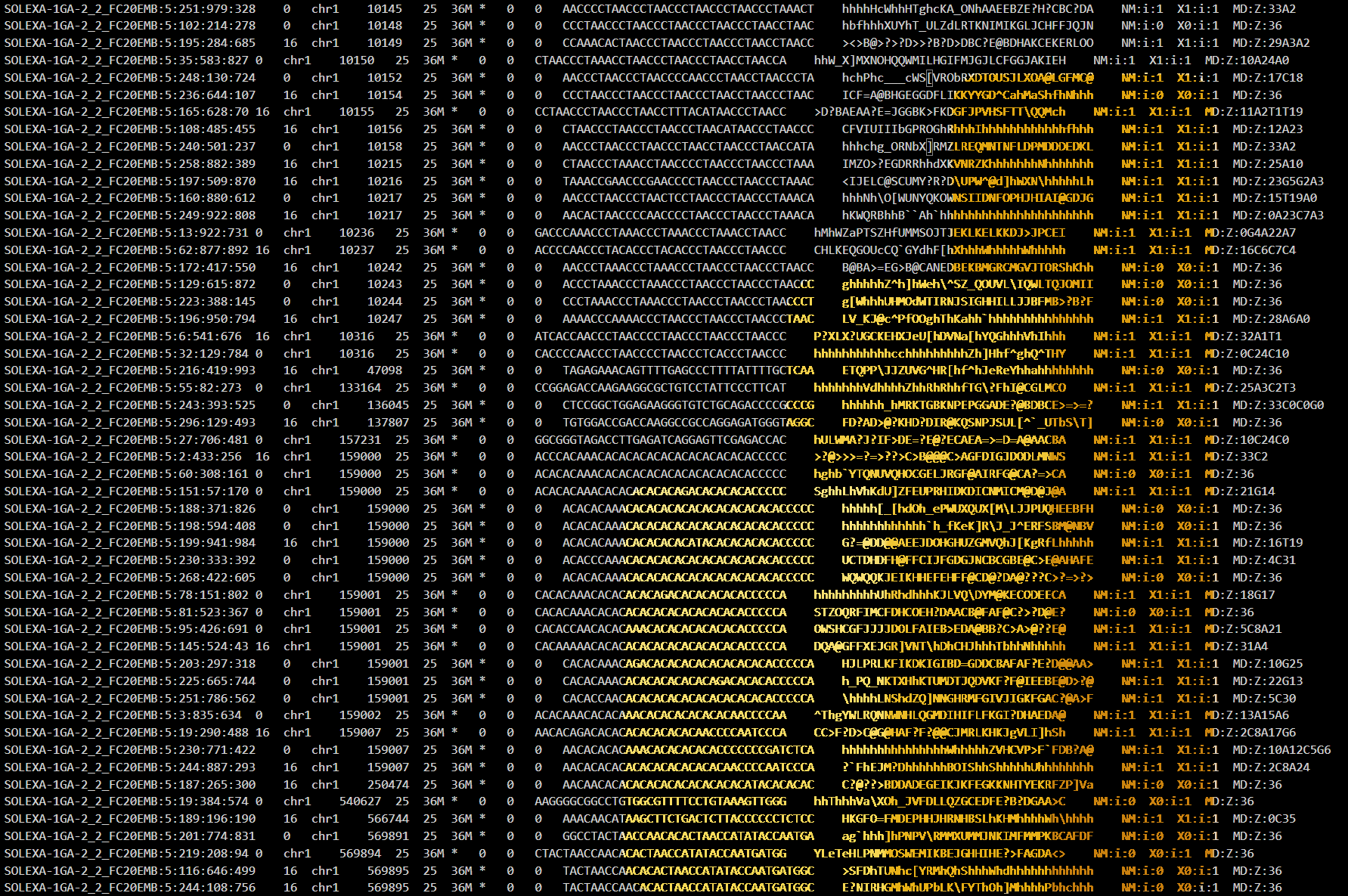
In the query editor, we can write a custom M script to parse our files. For example, to parse a SAM file:
|
|
Binary Files
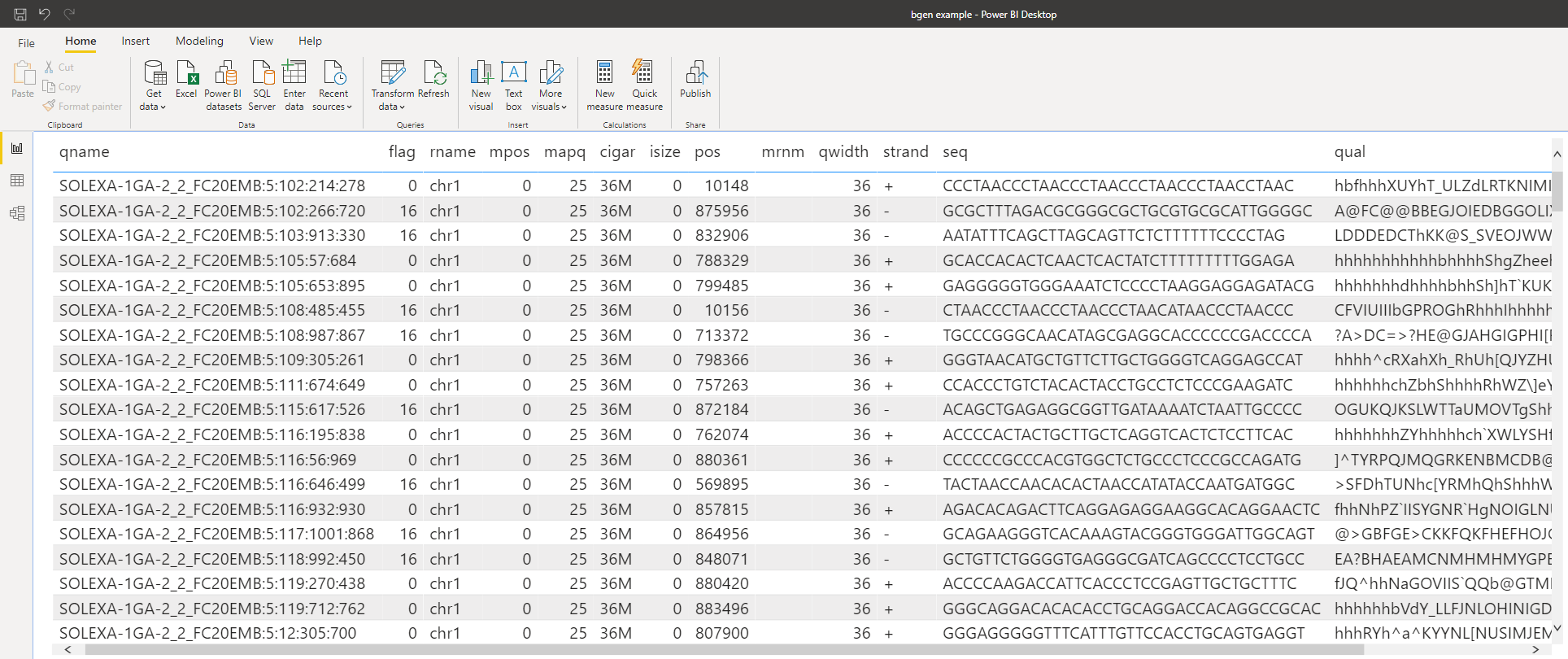
We can use R or Python to read in those pesky binary files as well. (This sometimes make even parsing text files simpler, too.) For example, to parse a BGEN file:
…which easily makes the file available as a table.
Online Sources
Lastly, Power BI makes it easy to grab data from the web. For example, let’s say that I wanted to get a list of all annotated genes of SARS-CoV-2 from NCBI.
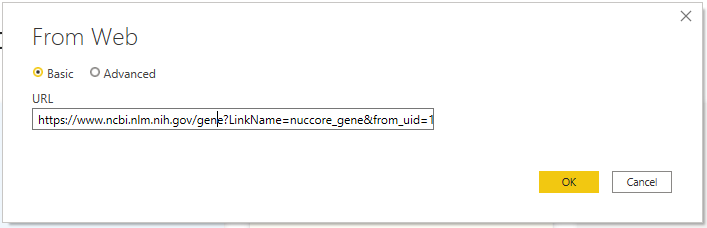
This enables users to take advantage of data from virtually any site. Try it out on the Protein Data Bank, NCBI, PlasmoDB, and more!
Takeaway Messages
- Be mindful of memory. Bioinformatics files can be large and, if you’re running on a machine with limited resources, you might bog it down.
- Check the defined specifications of any file format you’re looking to parse.
- R or Python can be your BFF, especially for binary or really complex file types.
Demo Video
Resources
All code used in above demos and additional examples are available at: www.github.com/BlueGranite/bioPowerBI
If you’d like to learn more about Power BI and how it can help you, contact 3Cloud today.